 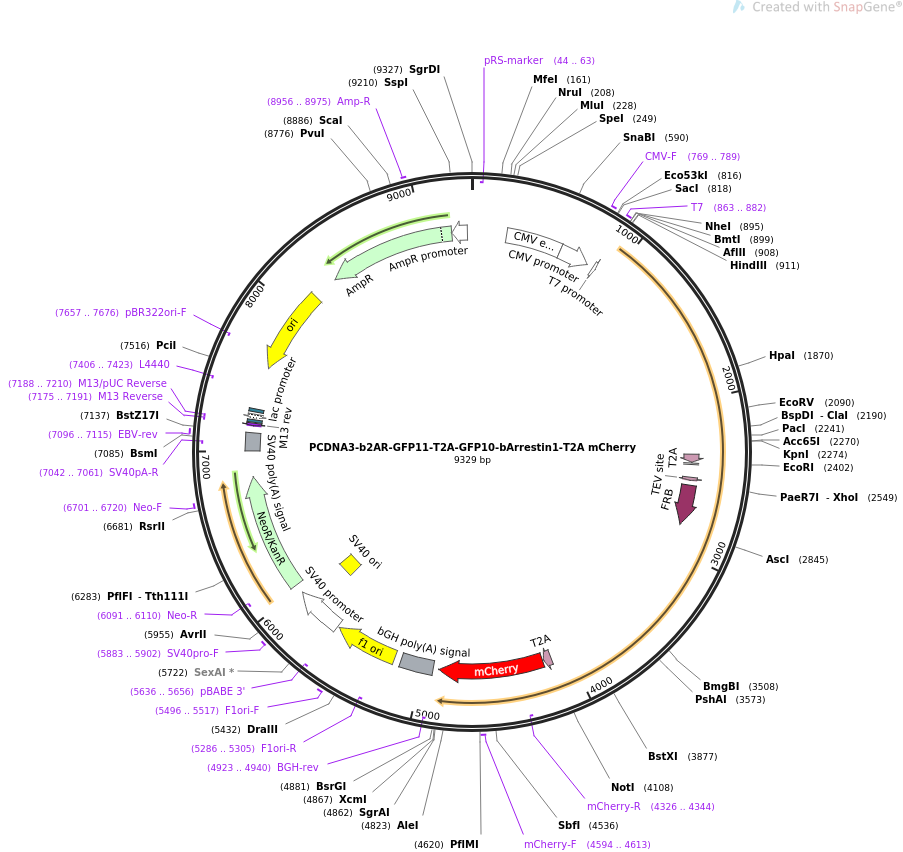
If two enzyme sites are too close together, one or both enzymes might not cut efficiently. The majority of enzymes require 4 to 6 base pairs adjacent to the recognition site to stabilize the DNA for efficient cleavage. The relative location of the enzymes to the end of your linear DNA molecule is also important. Examples include the directional cloning of a gene or a cDNA into a vector. Directional cloning guarantees that your inserted fragment is introduced in the orientation you choose and reduces background colonies resulting from a linear vector closing up on itself. Single restriction digests aim to isolate many different fragments of a given genomic DNA or cDNA.ĭual enzyme or directional cloning uses two different enzymes. A typical example of this is when cloning a genomic DNA library or a cDNA random cleavage library. This is used for experiments where the orientation of the DNA fragment to be cloned is not important. In single enzyme cloning, the vector and insert are cut with a single restriction enzyme. Single or dual restriction enzyme cloning strategies can be used depending on the cloning sites available and the importance of correct orientation of the insert. All blunt ends are compatible with each other. DNA with protruding or ‘sticky’ ends will join to compatible ends to form stable Watson-Crick base pairs. Once cleavage has occurred, the ends of the DNA become substrates for DNA ligase. Cloning Selection Marker: A marker that signals the vector contains inserted DNA.An MCS is a cluster of unique restriction enzymes that offer multiple enzyme choices for introducing your fragment of interest. Convenient Insertion Site: When cloning with restriction enzymes, the most common entry point for your fragment of interest is a Multi-Cloning Site or MCS.In bacteria, most selectable markers are antibiotic resistance genes. Selectable Marker: The selectable marker allows selective growth agents to guarantee that growing cells contain the vector.Origin of Replication: The origins of replication allow the cloned DNA to replicate independently of the genome.The plasmid below illustrates the essential features required of a restriction cloning vector. ► Watch an Introduction to Molecular Cloning video series Planning a Restriction Cloning Experiment Selecting a Vector System In fact, more than 70% of all molecular biology experiments begin with the restriction cloning of DNA fragments. Cloned genes are introduced into plants to improve their resistance to a pathogen or toxic chemical in their environment.Ĭloning and subcloning with restriction enzymes has been the dominant technique in molecular biology for many years.The CRISPR enzyme Cas-9 is cloned in many organism-specific vectors that allow CRISPR based genome editing to be applied in those organisms.Human insulin was cloned and expressed in bacteria in 1978.Genes are cloned so that the proteins they encode can be expressed at high levels and purified for medical, experimental or commercial applications. The development of these tools led to the publication of the first recombinant DNA molecules in 1972. The second major advance in the field was the development of plasmid cloning vectors that could be used to receive and replicate isolated pieces of DNA. This key discovery, coupled with the description of scientific protocols, enabled scientists to use these tools to isolate individual genes from a genome. The first advance was the discovery of restriction enzymes and the DNA ligase enzyme. Prior to the 1970s, scientists were not able to easily isolate and study individual genes. History and Applications of Restriction Cloning 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
